You will never have as much time as during your PhD to collect high-quality data. I didn’t realize it at that time, but detailed data you really know and understand is a lifetime companion. I used mine for its main purpose during my PhD, but also to test new methods when I needed to test those with data. In addition, it contributed to several synthesis papers, including an ongoing one led by someone at my lab right now. This makes at least five papers I used this data so far. As the data is openly available for a long time it was also used by several other synthesis papers. All this preamble is to encourage you to love your data!
When I collected the data I did most of the bees id’s myself (I had lots of help from experts such as Jordi Bosch, but in the end, it was me who ran through all the samples). This implies I identified several individuals to moprhospecies level, and I couldn’t put a name in all my pollinators. I would say this is typical for many ecological studies, and we always cross fingers this is good enough at the ecological community level. More than 10 years later I decided to properly identify by taxonomonists the full collection (which I managed to keep all this time while traveling through three countries!)
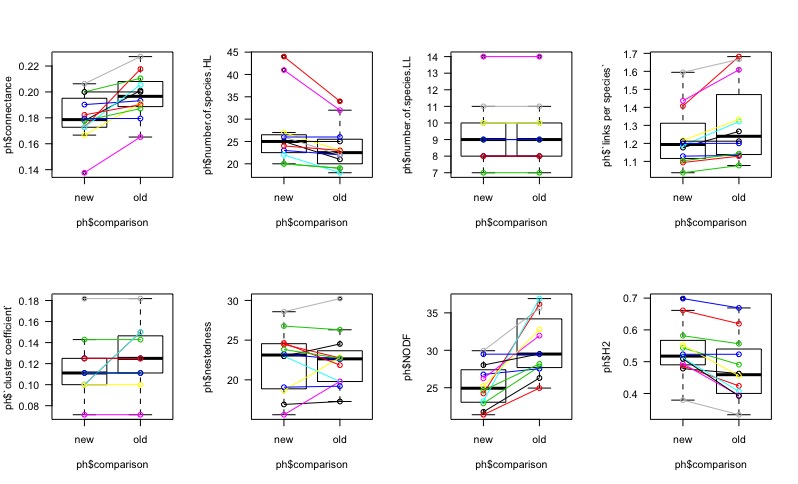
Almost 30% of individuals changed from morphospecies to being properly identified at the species level. Among those, most morphospecies belonged to a single species, but not all. In general, I underestimated the number of species present from 81 to 114. Several similar species were lumped together by my ignorance. However, to my relief, this did not change drastically the relative differences between the 12 sites, as seen in the figure. Connectance and the number of links per species decreased for all sites at similar rates, but some metrics are less consistent, such as nestedness, but nestedness is also a metric known to be quite volatile.
As I said, the data was released in different places. Web Interaction Database has a copy of the data in matrix format, which makes it hard to split the data e.g. by dates. Web of Life has even a more drastic pooling, which I am not sure how they did it, as I was not contacted by them, but I noticed all sites are pooled, including invaded and non invaded sites. FigShare had the best data so far, and it is associated with a paper and its analysis, so it’s better to keep this version at the morphospecies level for historical reasons. Hence, the new release of the data, with all new identifications is at Mangal. The webpage is very nice, and you can access programmatically all the networks in R.
library("rmangal")
mgs <- search_datasets("bartomeus")
mgn <- get_collection(mgs)
mgn; names(mgn)
mgn[[1]]$nodes
mgn[[1]]$interactions
library(tidygraph)
tg <- as_tbl_graph(mgn[[1]])
plot(tg)
I am a bit embarrassed by the lower quality of the original data, but better fixing it now than never. Long life to data!
